 Principles of timing mutations and copy number gains based on whole-genome sequencing
Principles of timing mutations and copy number gains based on whole-genome sequencingAbstract
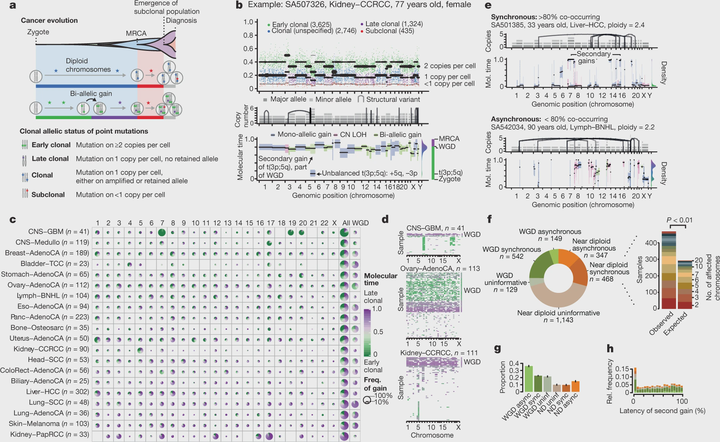
Cancer develops through a process of somatic evolution. Sequencing data from a single biopsy represent a snapshot of this process that can reveal the timing of specific genomic aberrations and the changing influence of mutational processes. Here, by whole-genome sequencing analysis of 2,658 cancers as part of the Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium of the International Cancer Genome Consortium (ICGC) and The Cancer Genome Atlas (TCGA), we reconstruct the life history and evolution of mutational processes and driver mutation sequences of 38 types of cancer. Early oncogenesis is characterized by mutations in a constrained set of driver genes, and specific copy number gains, such as trisomy 7 in glioblastoma and isochromosome 17q in medulloblastoma. The mutational spectrum changes significantly throughout tumour evolution in 40% of samples. A nearly fourfold diversification of driver genes and increased genomic instability are features of later stages. Copy number alterations often occur in mitotic crises, and lead to simultaneous gains of chromosomal segments. Timing analyses suggest that driver mutations often precede diagnosis by many years, if not decades. Together, these results determine the evolutionary trajectories of cancer, and highlight opportunities for early cancer detection.
More description of an article.